 Data Resources
Computational Analysis Tools
Standards
Data Hubs
Data Resources
Computational Analysis Tools
Standards
Data Hubs
| Name | McAN |
|---|---|
| Type | Phylogeny and Molecular Evolution |
| Version | V1.4.1 |
| Developers | Lun Li, Bo Xu, Anke Wang, Junwei Zhu |
| Description | McAN is a tool for constructing haplotype network based on minimum-cost arborescence. McAN overcomes the problems of slow processing speed and large memory consumption in traditional haplotype network construction algorithms, thereby enabling the evolutionary tracking analysis of massive SARS-CoV-2 sequences. Tests on a small dataset of around 1000 SARS-CoV-2 whole-genome data show that McAN runs two orders of magnitude faster than traditional methods. Tests on a large dataset of around 5 million SARS-CoV-2 sequences indicate that McAN can handle massive pathogenic genomic sequence data in about 25 minutes (with 50 threads). On test datasets, McAN has achieved accurate results while reducing memory consumption by over 90% compared with traditional methods. Moreover, McAN has demonstrated reasonable results on various datasets, including monkeypox and influenza A. |
| Downlaod | https://ngdc.cncb.ac.cn/biocode/tools/BT007301 |
| Online Link | https://ngdc.cncb.ac.cn/bit/hapnet/ |
| Article | https://doi.org/10.1093/bib/bbad174 |
| Figure |
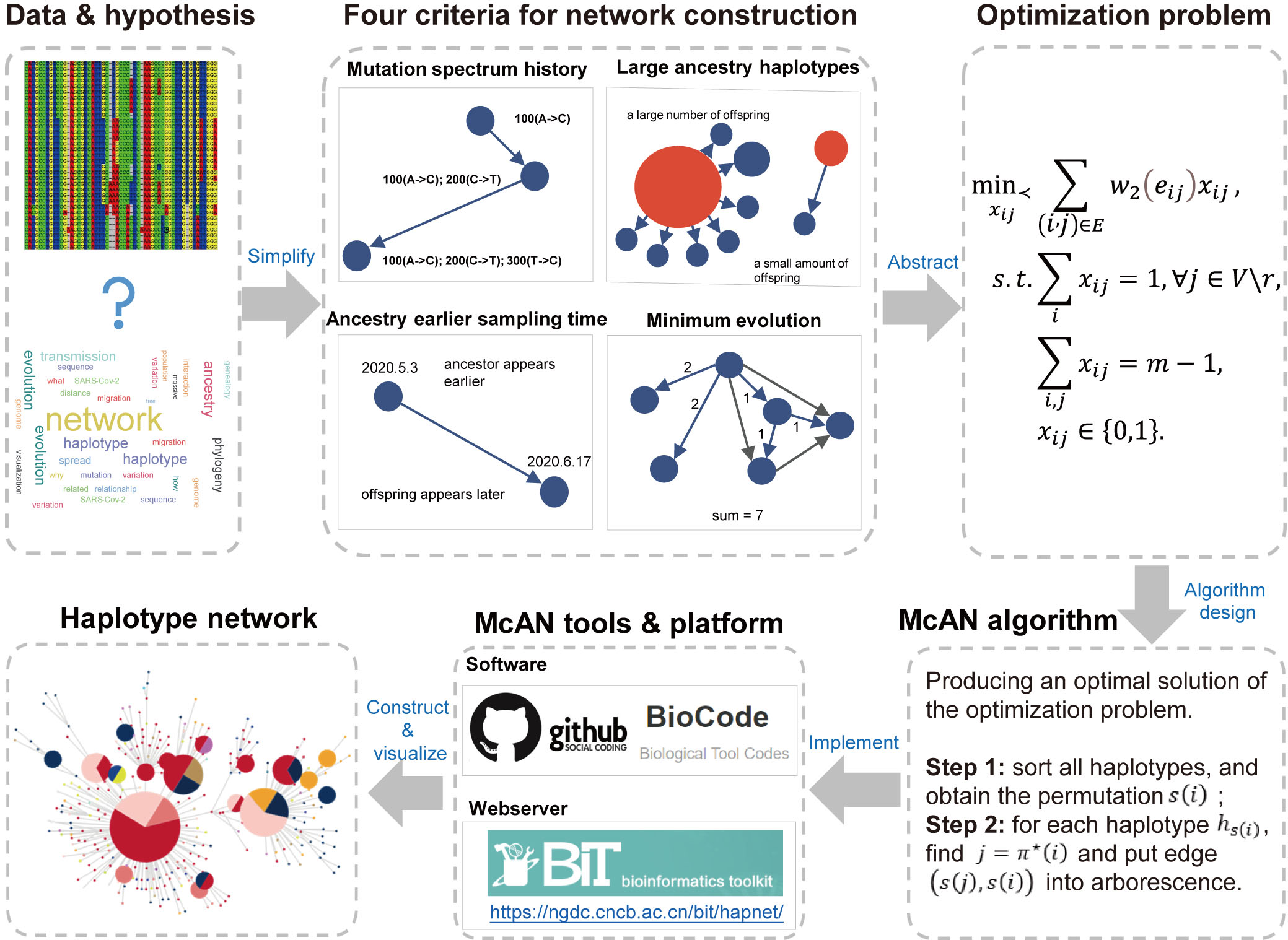
|

1 Beichen West Road, Chaoyang District Beijing 100101, China | 86-10-84097216
© China National Center for Bioinformation 2025, 京ICP备 10050270号-13